真菌数据的iCAMP计算,已经检查了导入数据行名列名都匹配还是一直报错,请问该如何解决:
报错如图:
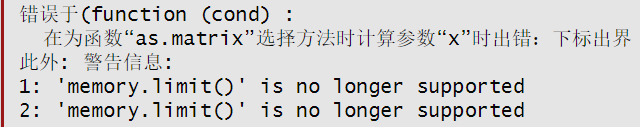
运行代码如下:
library(iCAMP)
library(picante)
library(ape)
rm(list=ls())
gc()
group <- read.csv('F_group.csv', row.names = 1)
group
classification <- read.csv('F_taxonomy_table_16S.csv', row.names = 1)
classification
tree <- read.tree('outsf.tre')
tree
otu <- read.csv('F筛选后的OTU(0.001%).csv', row.names = 1)
otu <- t(otu)
#修剪系统发育树以仅包含群落数据集中存在的物种或具有非缺失性状数据的物种
tree <- prune.sample(otu, tree)
tree
set.seed(123)
icamp.out <- icamp.big(comm = otu, tree = tree, pd.wd = getwd(), ses.cut = 1.96, rc.cut = 0.95, bin.size.limit = 12, rand = 1000, nworker = 10)
icampbin=icamp.bins(icamp.detail = icamp.out, treat = group,
clas = classification, boot = TRUE,
rand.time = 1000, between.group = TRUE)
#输出
write.csv(icamp.out$CbMPDiCBraya, 'icamp.out.bacteria.csv', row.names = FALSE)
write.csv(icampbin$Bin.Topclass, 'Bin.topclass.csv', row.names = FALSE)
write.csv(icampbin$Class.Bin, 'Bin.composition.csv', row.names = FALSE)
write.csv(icampbin$BPtk, 'Bin.contribution.to.process_eachgroup.csv', row.names = FALSE)
write.csv(icampbin$Ptk, 'Bin.process.relative.importance.csv', row.names = FALSE)
