import os
import numpy as np
from matplotlib import pyplot as plt
from tensorflow.keras.layers import Conv2D, BatchNormalization, Activation, MaxPool2D, Dropout, Flatten, Dense
from tensorflow.keras import Model
np.set_printoptions(threshold=np.inf)
cifar10 = tf.keras.datasets.cifar10
(x_train, y_train), (x_test, y_test) = cifar10.load_data()
x_train, x_test = x_train / 255.0, x_test / 255.0
class Baseline(Model):
def __init__(self):
super(Baseline, self).__init__()
self.c1 = Conv2D(filters=6, kernel_size=(5, 5), padding='same') # 卷积层
self.b1 = BatchNormalization() # BN层
self.a1 = Activation('relu') # 激活层
self.p1 = MaxPool2D(pool_size=(2, 2), strides=2, padding='same') # 池化层
self.d1 = Dropout(0.2) # dropout层
self.flatten = Flatten()
self.f1 = Dense(128, activation='relu')
self.d2 = Dropout(0.2)
self.f2 = Dense(10, activation='softmax')
def call(self, x):
x = self.c1(x)
x = self.b1(x)
x = self.a1(x)
x = self.p1(x)
x = self.d1(x)
x = self.flatten(x)
x = self.f1(x)
x = self.d2(x)
y = self.f2(x)
return y
model = Baseline()
model.compile(optimizer='adam',
loss=tf.keras.losses.SparseCategoricalCrossentropy(from_logits=False),
metrics=['sparse_categorical_accuracy'])
checkpoint_save_path = "./checkpoint/Baseline.ckpt"
if os.path.exists(checkpoint_save_path + '.index'):
print('-------------load the model-----------------')
model.load_weights(checkpoint_save_path)
cp_callback = tf.keras.callbacks.ModelCheckpoint(filepath=checkpoint_save_path,
save_weights_only=True,
save_best_only=True)
history = model.fit(x_train, y_train, batch_size=16, epochs=5, validation_data=(x_test, y_test), validation_freq=1,
callbacks=[cp_callback])
model.summary()
# print(model.trainable_variables)
file = open('./weights.txt', 'w')
for v in model.trainable_variables:
file.write(str(v.name) + '\n')
file.write(str(v.shape) + '\n')
file.write(str(v.numpy()) + '\n')
file.close()
############################################### show ###############################################
# 显示训练集和验证集的acc和loss曲线
acc = history.history['sparse_categorical_accuracy']
val_acc = history.history['val_sparse_categorical_accuracy']
loss = history.history['loss']
val_loss = history.history['val_loss']
plt.subplot(1, 2, 1)
plt.plot(acc, label='Training Accuracy')
plt.plot(val_acc, label='Validation Accuracy')
plt.title('Training and Validation Accuracy')
plt.legend()
plt.subplot(1, 2, 2)
plt.plot(loss, label='Training Loss')
plt.plot(val_loss, label='Validation Loss')
plt.title('Training and Validation Loss')
plt.legend()
plt.show()
==============================================================================================
以上是相关代码,课程中 正确率结果约为50%, 为什么我用的是一样的代码 跑出来是0.1
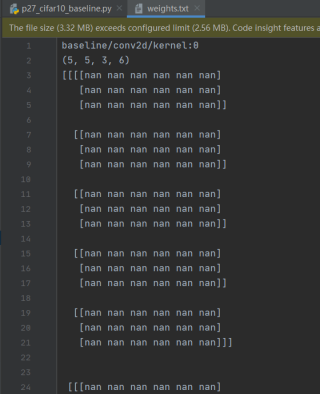
下面是weights.txt文件, 发现卷积核的参数全是nan